Building New Matrix
To create a position weight or frequency matrix from an alignment or a file with several sequences, press the Build new matrix button in the Weight matrix search dialog, or select the Tools ‣ Search for TFBS ‣ Build weight matrix main menu item:
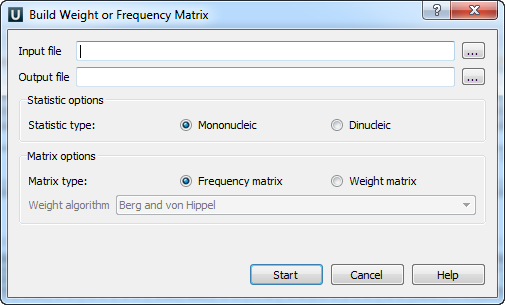
The Build weight or frequency matrix dialog will appear:
The following parameters are available:
Input file — an alignment or a file with several sequences to build the matrix from. This parameter is mandatory.
Output file — the resulting matrix will be saved in this file. This parameter is mandatory.
Statistic type — defines how the statistics will be collected. The Mononucleic option is generally good for small alignments, while the Dinucleic option should give more appropriate results for large alignments.
Matrix type — defines the type of the resulting matrix.
- If the Frequency matrix option is selected, then the frequency matrix will be created and saved in the resulting file.
- If the Weight matrix option is selected, then an intermediate frequency matrix will be created and then transformed into a weight matrix based on the selected Weight algorithm. The weight matrix will then be saved into the resulting file.
For some input files, the colored “Alignment Logo” appears at the bottom of the dialog. It provides a representation of the selected alignment.
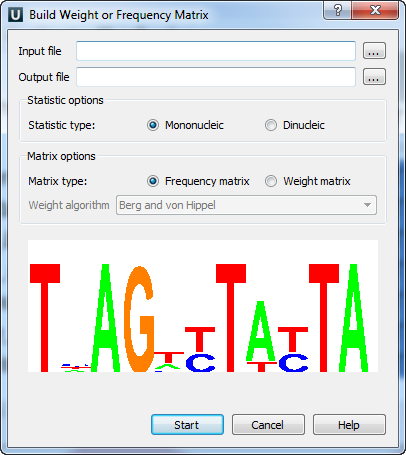
The “Alignment Logo” appears when:
- The input file format is *.pfm, *.aln, or it is a file with several sequences.
- The size of the input file is small enough.
To start the operation, press the Start button. The matrix will be created and saved. If the Build weight or frequency matrix dialog was invoked from the Weight matrix search dialog, then the matrix will also be chosen as the current profile.