Creating Annotation
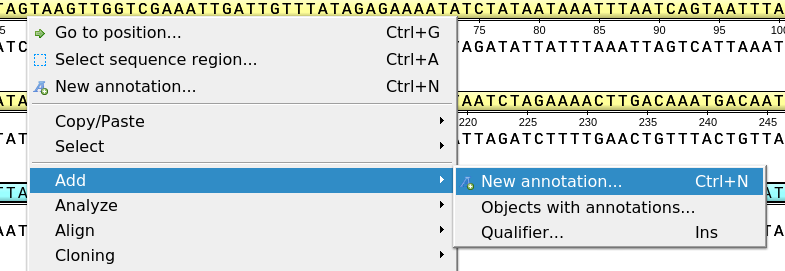
To create a new annotation for the active sequence, press the Ctrl-N key sequence, select the New annotation toolbar button, or use the Add ‣ New annotation or New annotation context menu item:
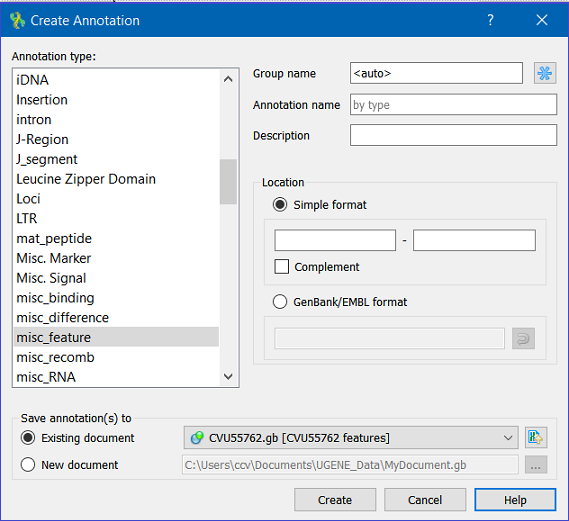
This will activate a dialog to set up annotation parameters:
The dialog asks where to save the annotation. It could be either an existing annotation table object or a new annotation table.
You can also specify the name of the group and the name of the annotation. If the group name is set to
The Location field contains annotation coordinates. The coordinates must be provided in the Genbank or EMBL file formats. If you want to annotate a complement strand sequence, check the corresponding checkbox for the simple format, or surround the coordinates with the “complement()” word, or press the last button in the corresponding row to do it automatically.
Note that by default the Location field contains the coordinates of the selected sequence region.
For amino acid sequences, there is no “Complement” checkbox in the “Simple format” group in the Location section, and there is no “Add/remove complement flag” button in the “GenBank/EMBL format” group.
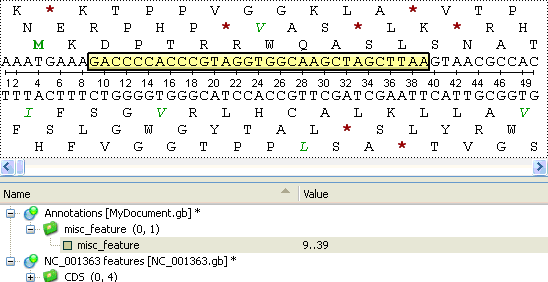
Once the Create button is pressed, the annotation is created and highlighted both in the Sequence overview and the Sequence details view areas: