Filter Sequence That Matches a Pattern
This workflow allows filtering sequences that match (or do not match) user-specified patterns.
How to Use This Sample
If you haven’t used workflow samples in UGENE before, refer to the “How to Use Sample Workflows” section of the documentation.
Workflow Sample Location
The workflow sample “Filter Sequence That Match a Pattern” is available in the Scenarios section of the Workflow Designer samples.
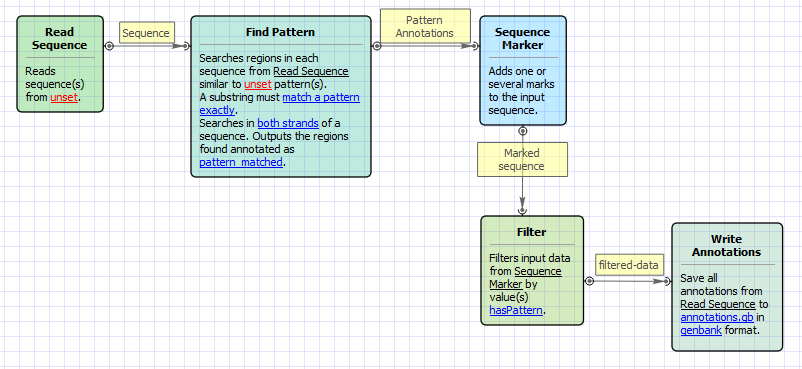
Workflow Image
The workflow looks as follows:
Workflow Wizard
The wizard consists of 3 pages:
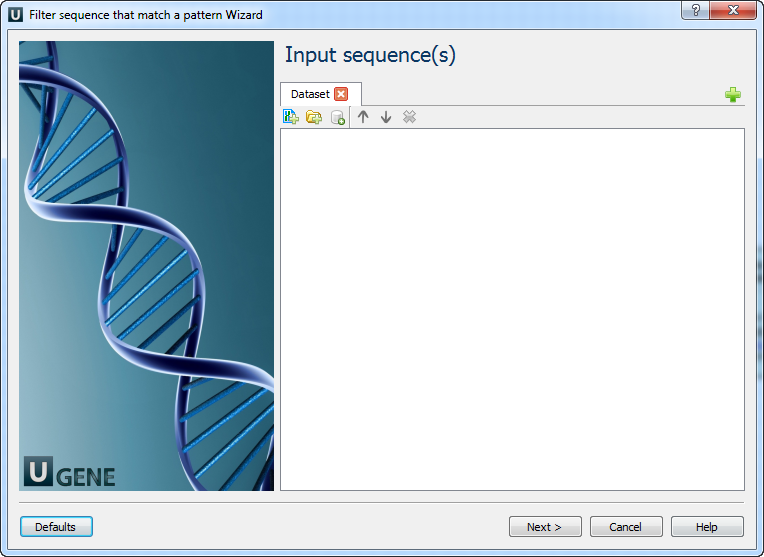
1. Input sequence(s)
On this page, input sequence files must be selected.
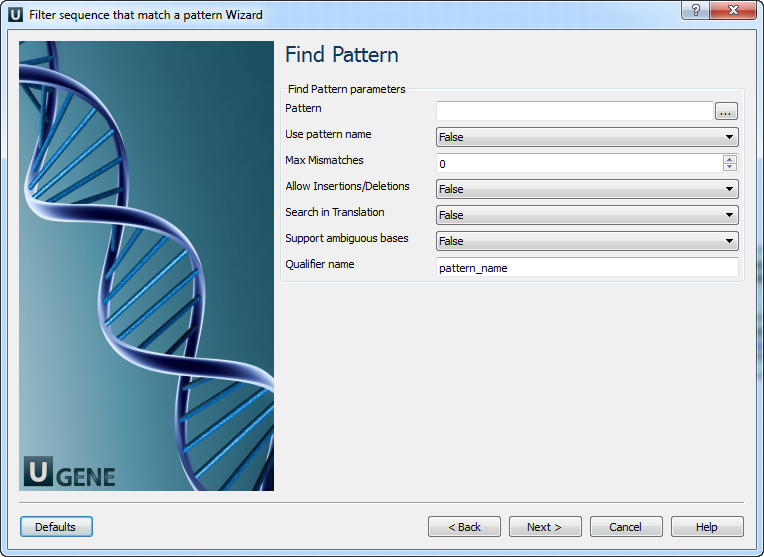
2. Find pattern
On this page, patterns must be specified, and search parameters can be configured.
Parameters
| Parameter | Description | Default Value | Type |
|---|---|---|---|
| Pattern | Semicolon-separated list of patterns to search for | (user-defined) | string |
| Use pattern name | Use names of pattern sequences as annotation names (if loaded from file) | False | boolean |
| Max Mismatches | Maximum number of mismatches allowed between a substring and a pattern | 0 | numeric |
| Allow Insertions/Deletions | Consider insertions/deletions in search | False | boolean |
| Search in Translation | Translate nucleotide sequence into protein and search in translated sequence | False | boolean |
| Support ambiguous bases | Handle ambiguous bases correctly (disables insertions/deletions when enabled) | False | boolean |
| Qualifier name | Name of the qualifier in result annotations containing the pattern name | pattern | string |
3. Output data
This page allows configuring the output file.