Overview of NGS Pipelines in UGENE
UGENE is a free bioinformatics platform that integrates dozens of well-known biological tools and algorithms and provides both graphical and command line interfaces for them. It is designed to solve various complex computational tasks in bioinformatics. One of the areas is analysis of sequencing data produced by popular modern NGS technologies.
There are three basic pipelines integrated into UGENE for NGS data analysis: Variant Calling, RNA-sequencing data analysis and ChIP-sequencing data analysis. The RNA-seq and ChIPseq (chromatin immunoprecipitation sequencing) pipelines have more than one possible configurations.
Variant Calling with SAMtools
The Variant Calling pipeline uses the SAMtools software to identify genomic variants in the assembled sequencing data by comparing them with a reference sequence.
Tuxedo – RNA sequencing protocol
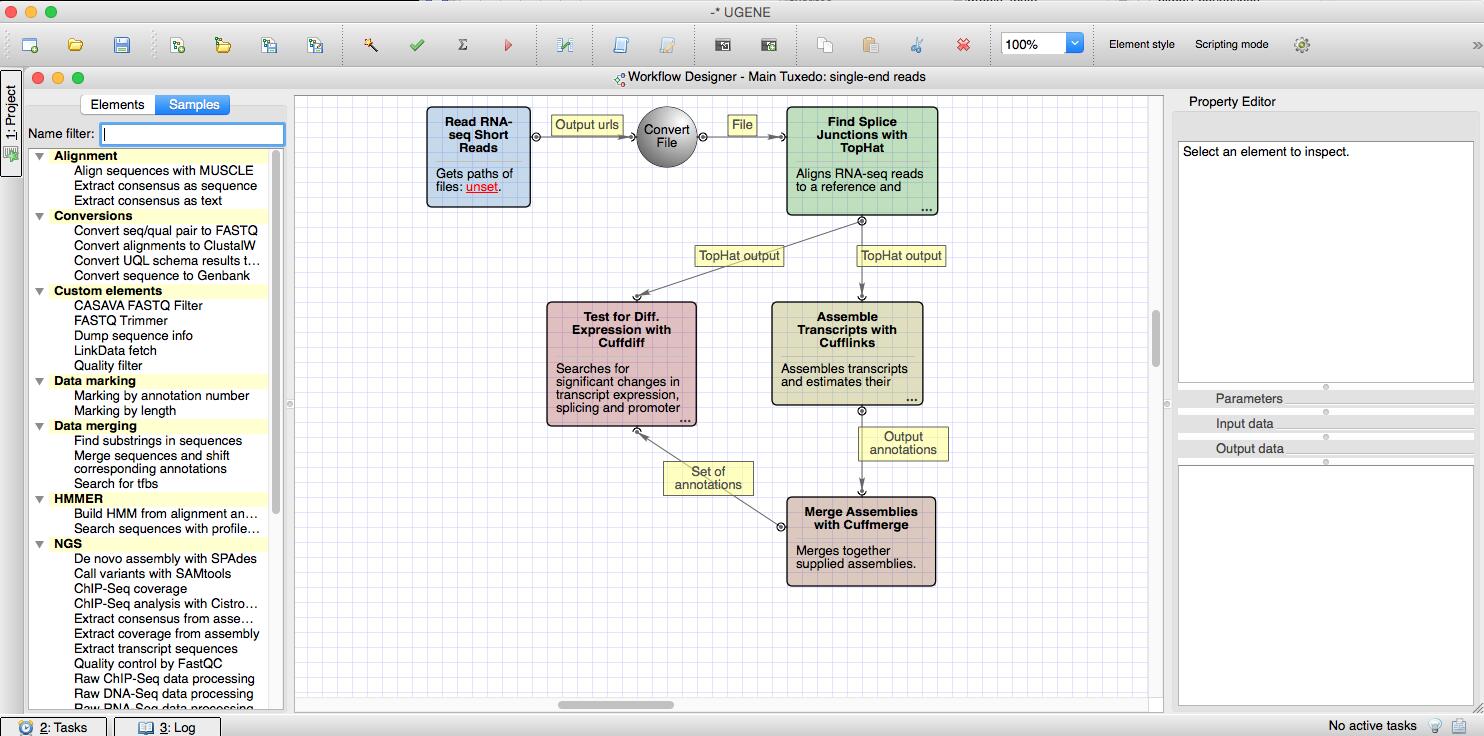
The RNA sequencing protocol is based on the Tuxedo toolkit and contains TopHat and the Cufflinks RNA tools. The pipeline helps to discover new genes, new splice variants of known genes (Cufflinks RNA) and to compare genes and transcripts expression in RNA-seq experiments.
Cistrome
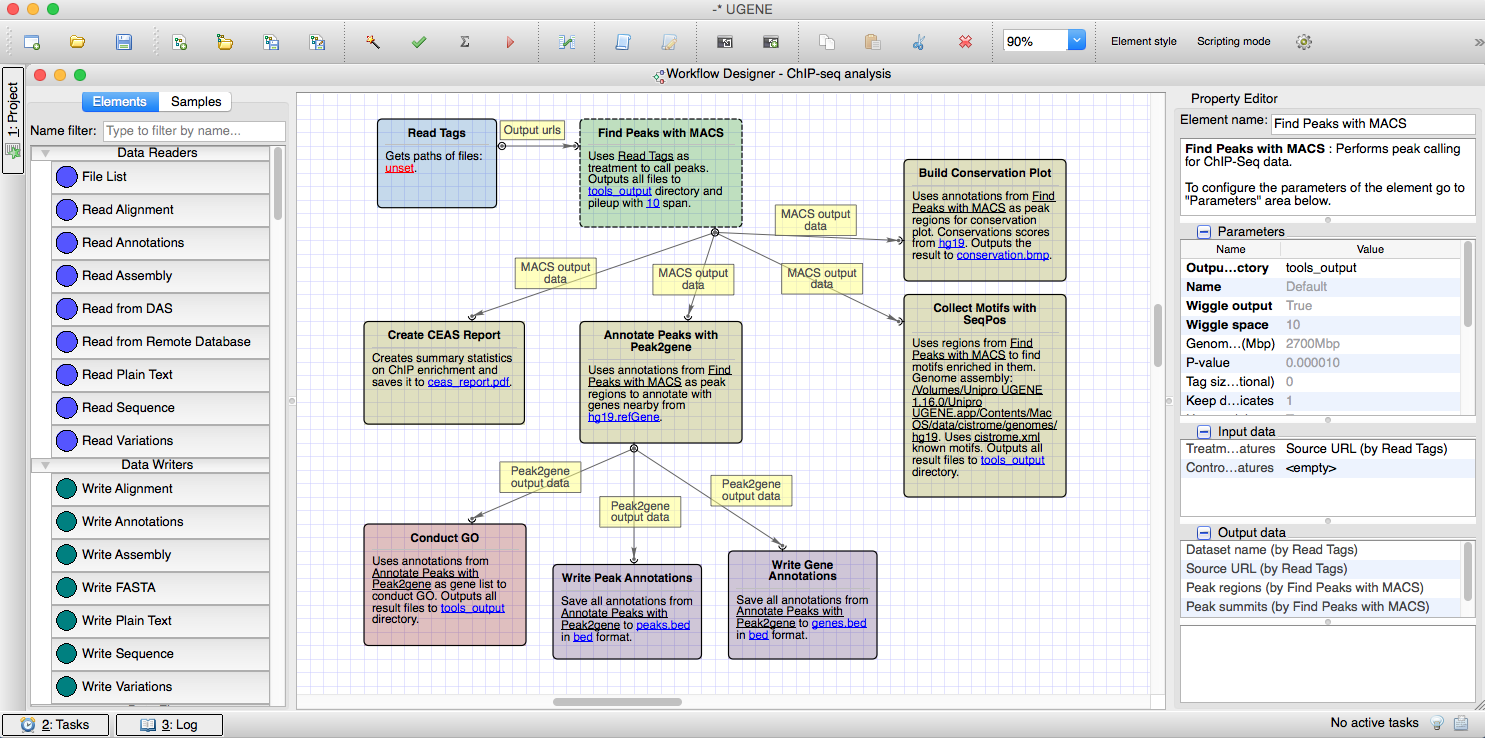
The ChIPseq pipeline is based on the tools from the Cistrome platform. The tools together provide a way of gene expression analysis and motif discovery with preliminary peak calling in chromatin immunoprecipitation sequencing experiments.
Workflow Designer
All the pipelines can be used in the UGENE Workflow Designer – the graphical system for construction of the computational workflows. A user with a multi-stage computational task can use the Workflow Designer and easily create a workflow that would solve the task. Once created, the workflow can be used multiple times for performing calculations on different input data.
There are ready-to-use samples in the Workflow Designer. The NGS pipelines are located in the “NGS” section of the samples. By double-clicking on a sample and supplying the input data you can run a pipeline with default parameters and outputs.
Using the NGS pipelines in UGENE a user have a set of advantages over using the original tools. Configurations of the Tuxedo and the Cistrome pipelines can be selected in the special dialog before running the workflow.
Each NGS workflow is equipped with wizards. A wizard is a simplified visual interface used to set the workflows most important parameters. The wizard is logically divided into pages and each page has its own set of settings. Going over the wizard a user sets the workflow parameters step by step and has detailed explanation for all of them.
Handy monitoring
After the workflow is configured and run a user is able to monitor the process using a dashboard. The dashboard provides the details about the status of the running process, progress, files produced and so on. Advanced users can see logs of the external tools in a special section of the dashboard.
Results managing
Dashboards persist between launches of the UGENE. So a user can see the information about previous runs, check parameters of a run and open the produced files.
Even a closed dashboard could be restored using the Dashboards Manager. If a user does not want to keep a dashboard it can be removed it with all data produced using the manager.
Availability
To run basic pipelines user should download UGENE NGS package from the UGENE web-site. The basic pipelines are available out-of-the box and do not require additional configuration. These pipelines can work in offline mode using the Internet connection only for extra features.
UGENE makes NGS data analysis easier. Try it yourself!

