Tutorial: Building Phylogenetic Trees in UGENE
This tutorial is about phylogenetic trees building methods available in the UGENE platform. So here UGENE is presented as a phylogenetic tree maker.
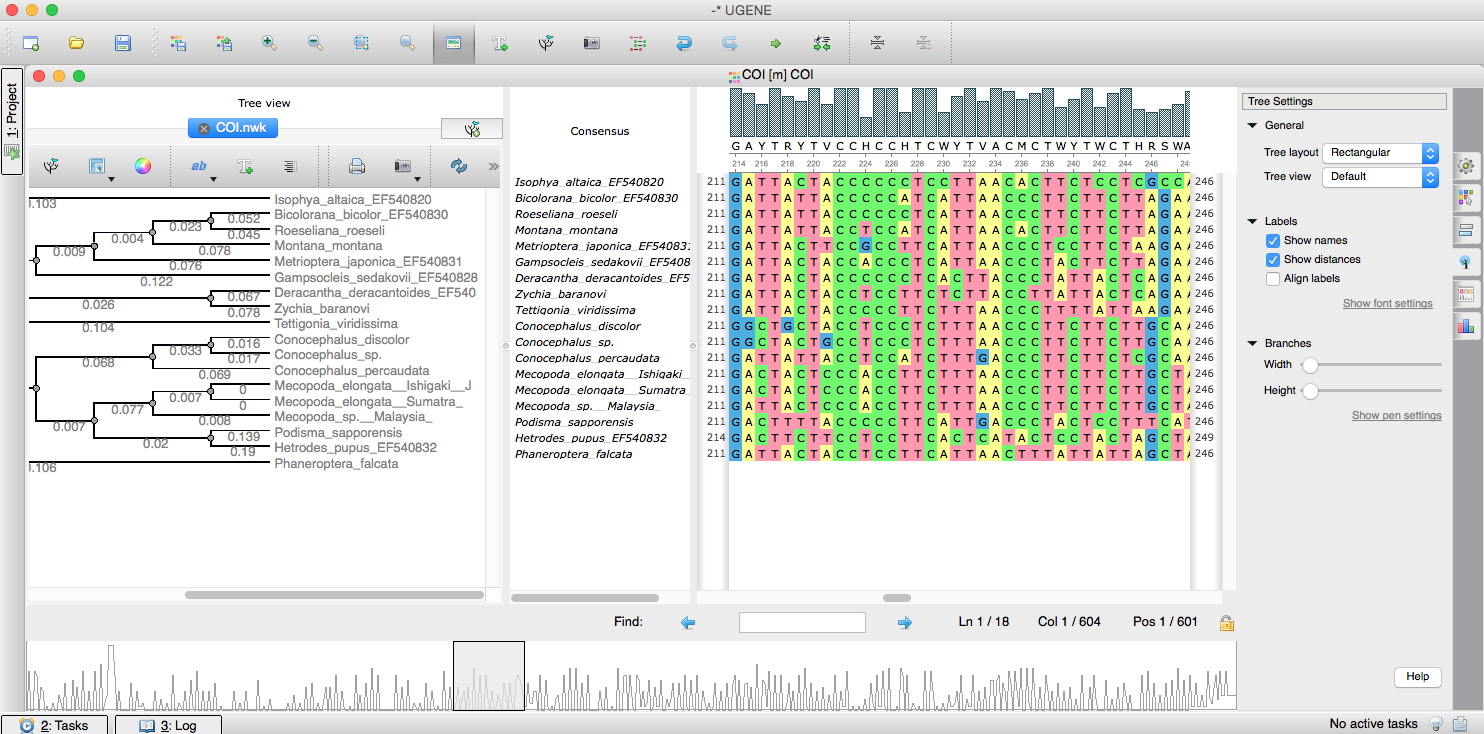
UGENE allows you to visualize, edit and build phylogenetic trees and phylogenetic relationship. You can also connect it to your multiple alignment and edit the tree with its sequences.
Phylogenetic Tree Methods Overview
There are three different phylogenetic trees building methods based on different algorithms: Neighbor-joining from the PHYLIP toolset, Bayesian inference from the MrBayes software and maximum likelihood from PhyML. Each method has its own strong and weak features related to the acceptable sizes of input alignments, required computational resources, etc. All these methods, integrated into UGENE, require the multiple sequence alignment file as an input. Let’s consider them one by one.
You might want to check PHYLIP -> PHYLIP on the web -> may be read here -> Neighbor-Joining and UPGMA method documentation files.
The neighbor joining method is a distance-matrix method. It is the commonly used method because it is simple, straightforward and fast, and usually it can be run with the default settings. But the method is suitable only for small multiple alignments.
Running PHYLIP
Let’s perform the test run of PHYLIP Neighbor-Joining. Open the sample multiple alignment in UGENE. Click the “Build tree” button on the main toolbar and choose the needed method. Run the building process. The result will be automatically opened in the UGENE tree viewer.
Running PhyML
The PhyML maximum likelihood method was designed to process large data sets. In theory, it can process alignments with up to 4,000 sequences 2,000,000 character-long. PhyML has the large number of substitution models and settings. It makes possible to build phylogenetic trees going from very fast and efficient methods (for a simple phylogenetic tree) to slower but more accurate ones.
For running PhyML, repeat the previous scenario with another sample file but choose the maximum likelihood method in the dialog.
Running MrBayes
And the third method is the Bayesian inference from the MrBayes software. The method is based on building a set of possible phylogenetic trees and assuming a prior probability distribution of each tree. MrBayes is more computational resources consuming than PHYLIP neighbor joining but less than PhyML phylogenetic tree maker.
Let’s run the method using the MrBayes sample. Open the file in UGENE, click the familiar “Build tree” button and choose the MrBayes method. [Pause] Choose the GTR substitution model and set the Rate option as invgamma like it is made in the MrBayes tutorial.
Conclusion
UGENE integrates the algorithms making them easier to use keeping you apart of command line interface and operations on file formats.
Note that these are only the methods of building phylogenetic relationship, you might want to check visualization and simple phylogenetic tree editing options in documentations. UGENE also support multiple formats of files with trees.

