Tutorial: How to Use Primer3 to Design Primers in UGENE
UGENE Primer3 primer design plugin allow designing primers. You can suggest Polymerase Chain Reaction primers to create sequence tagged sites for hybrid mapping, or to amplify sequences for single nucleotide polymorphism discovery, and for other applications.
The Primer3 primer design plugin is based on the widely used open-source Primer3 tool (developed by Steve Rozen and Helen Skaletsky). Its interface is similar to the web-interface of Primer3, therefore if you have used the WWW program version, it won't take much effort for you to use the UGENE version.
Setting Primer Design Parameters
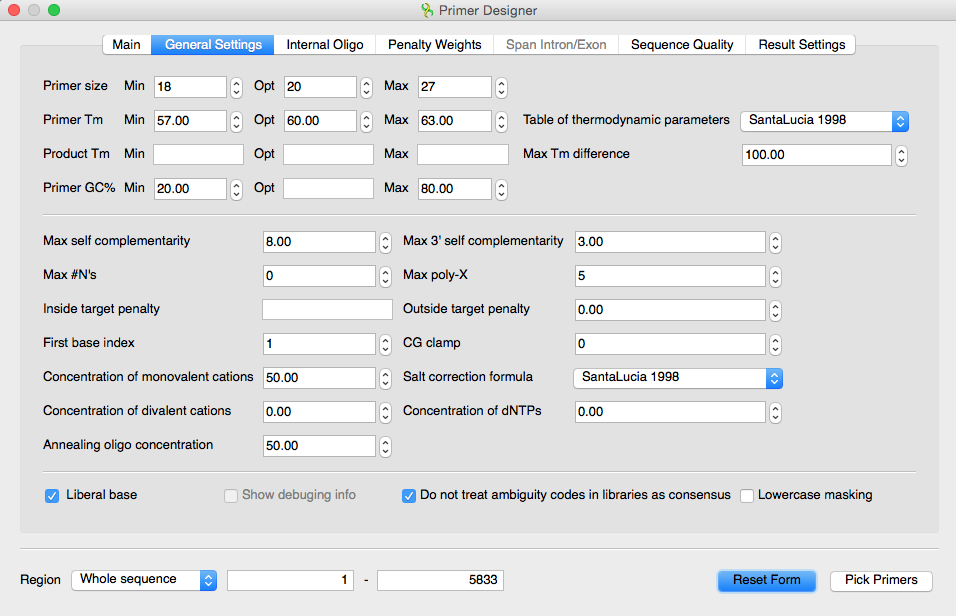
I have UGENE with an opened sequence. To invoke the plugin, click the „Primer3“ button, or right-click on a sequence view and select „Analyze → Primer3“. The „Primer Design“ dialog will be opened to design primers. It has plenty of options, just like the web interface does, but eventually it allows you to do sort of "virtual PCR". Thus you can describe the variety of designing primers criteria, including such factors as oligonucleotide melting temperature, size, GC content, PCR product size, positional constraints within the sourse sequence and many more.
I, on the other hand, will run a very simple "virtual PCR" example, to show you how the plugin works. Most of the default settings of the plugin are strict. So, I will specify a target. A Target might be a simple sequence repeat (microsatellite) site (for example a CA repeat) or a single-base-pair polymorphism. I will specify it with the start value of 40 (which is the index of the first base of a Target), and of the length of 78. A legal primer pair must flank this target since it's the only one specified.
I will also want Primer3 to pick internal oligo along with left and right primers.
The resulting primers are saved and shown as annotations. The „Result Settings“ tab allows to specify the annotations table to contain the results. I will call the results group „primers. Now I press „Pick Primers“.
Saving Primers
The primers for the selected sequence are found. The results annotations group contains a subgroup for every found primer pair.
This is how you can design primers in UGENE.

